Introduction
The path of a biopharmaceutical from research and development to consumer availability is a long and thorough process, intended to ensure efficacious and quality control. Development of biosimilars requires evaluation of the molecule’s HOS (higher order structure) with the original biopharmaceutical1). Additionally, the structure of biopharmaceuticals is sensitive to changes in the manufacturing process or storage environment and it is necessary to evaluate the HOS of these molecules sustained after any processing or storage alterations2).
CD spectroscopy is widely used to assess HOS in biopharmaceuticals because the far-UV CD is sensitive to changes in the bond angles of a protein’s backbone chain and secondary structure and the near-UV monitors the tertiary structure, composed of aromatic amino acids residues and disulfide bonds. Here, we evaluated the HOS of three types of novel anti-coronavirus (SARS-CoV-2) VHH antibodies (variable domain of heavy chain of heavy chain antibody) using the [Spectrum QC Compare Test] program that quantitatively determines the significance in structure changes based on the difference in CD spectral shape.
CD Spectra Measurement
– Far-UV CD spectrum
The CD spectrum in the far-ultraviolet region with shorter wavelength than 250 nm has absorption of the peptide bond, which is the main chain structure of the protein, and has a secondary structure (α-helix, β-sheet, turn, disordered structure). CD spectra in the far-ultraviolet region were measured in this application in order to evaluate the identity of the secondary structure.
– Near-UV CD spectrum
The CD spectrum in the near-ultraviolet region of 250 to 300 nm is attributed to the absorption related with aromatic amino acid side chains of proteins and the absorption of disulfide bonds. It is used to analyze interactions. Here, CD spectra in the near-ultraviolet region were measured to evaluate the identity of the tertiary structure as structure of the surrounding environment of tryptophan, tyrosine, and phenylalanine and interactions between aromatic amino acid.
Determination of differences in CD spectra
(1) Calculation of correlation coefficient: The CD spectrum of the reference sample is measured multiple times, and the correlation coefficient between the average spectrum (R(λ)) and the CD spectrum U(λ) of the sample is calculated.
(2) Weighting: The [Spectrum QC Compare Test] program allows the user to choose whether to weight the spectrum. The CD spectrometer has a low S/N due to weak light energy detected in the shorter wavelength region as well as the wavelength region where sample absorption is strong. Therefore, it is possible to reduce the weighting of the spectrum in the wavelength range where the effects from noise are large and increase the weighting in the wavelength range where noise is small (U.S. Patent 10,564,035. 2020-02-18). As a result of this data processing, the influence of noise can be reduced in the wavelength range where the S/N is smaller, and the spectral difference between samples can be determined with high sensitivity.
(3) Quantify the difference (i.e. distance (I)) between the CD spectra in between reference and sample.
(4) Judgment: The Z-test determines if there is a significant difference between the reference and sample spectra. If the Z-score is 2 or less, the reference and the sample are judged to be the same, and if the Z-score is larger than 2, the reference and the sample are judged to be different.
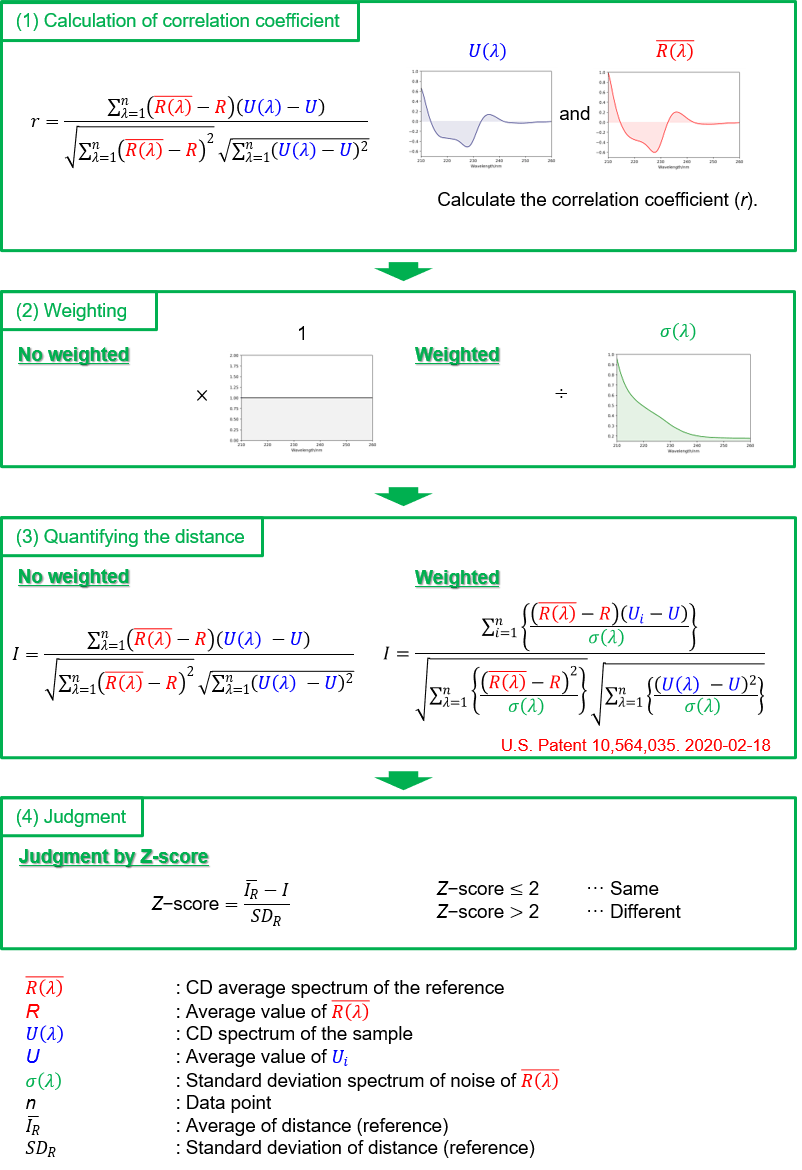
Figure 1. Analysis steps for determining the spectral differences between the reference and the sample spectra.
Experimental
Sample
– CD spectrum measurement (Far-UV region)
anti-SARS-CoV-2 VHH antibody (3 kinds) : No.1 (0.2 mg/mL), No.2 (0.2 mg/mL), No.3 (0.2 mg/mL)
Buffer : 20 mM PBS
– CD spectrum measurement (Near-UV region)
anti-SARS-CoV-2 VHH antibody (3 kinds) : No.1 (6.27 mg/mL), No.2 (7.22 mg/mL), No.3 (5.15 mg/mL)
Buffer : 20 mM PBS
(All VHH antibody samples were provided by RePHAGEN 3). )
Measurement parameters
– CD spectrum measurement (Far-UV region)
Wavelength range : 250 – 200 nm
Bandwidth : 1 nm
D.I.T.* : 4 sec.
Scanning speed : 50 nm/min.
Data interval : 0.1 nm
Accumulations : 1
– CD spectrum measurement (Near-UV region)
Wavelength range : 310 – 250 nm
Bandwidth : 1 nm
D.I.T.* : 4 sec.
Scanning speed : 20 nm/min.
Data interval : 0.1 nm
Accumulations : 4
*D.I.T.: Time Integration Time

J-1500 CD Spectrometer
Keywords
Novel coronavirus, SARS-CoV-2, COVID-19, Biopharmaceuticals, Antibody drug, BioSimilar, VHH antibody, HOS, Secondary Structure, Tertiary Structure, Circular Dichroism
Results
Visual comparison of CD spectra
Figure 2 shows the CD spectra of VHH antibodies in the far-UV region and near-UV region. In the far-UV region, a negative signal at 217 nm characteristic of β-sheet structure was observed only for sample No. 2 and no significant difference was observed in the CD spectra for samples No. 1 and No. 3. In the near-UV CD spectra, signals derived from phenylalanine (Phe) at 255 to 270 nm, tyrosine (Tyr) at 275 to 282 nm, and tryptophan (Trp) at 290 to 305 nm were observed in all samples4). While the CD spectra of samples No. 2 and No. 3 are visually similar, the spectra of sample No. 1 is clearly different.
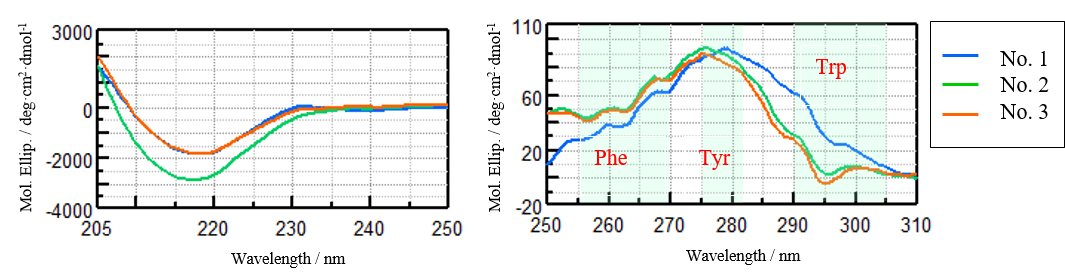
Figure 2. Far-UV (left) and near-UV (right) CD spectra of VHH antibodies for samples No. 1, 2, and 3.
Judgment of slight CD spectra difference by statistical method
The differences in the far-UV CD spectra of samples No. 1 and No. 3 and the near-UV CD spectra of samples No. 2 and No. 3 cannot be clearly discriminated. However, using the [Spectrum QC Compare Test] program, the distance between the spectra was obtained by the analysis method shown in Figure 1, and the differences judged by the Z-test. The number of spectral data used for the analysis is as follows and the individual spectra are shown in Figure 3:
Far-UV region: Reference No. 1 (9 spectra), sample No. 3 (5 spectra)
Near-UV region: Reference No. 2 (9 spectra), sample No. 3 (5 spectra)
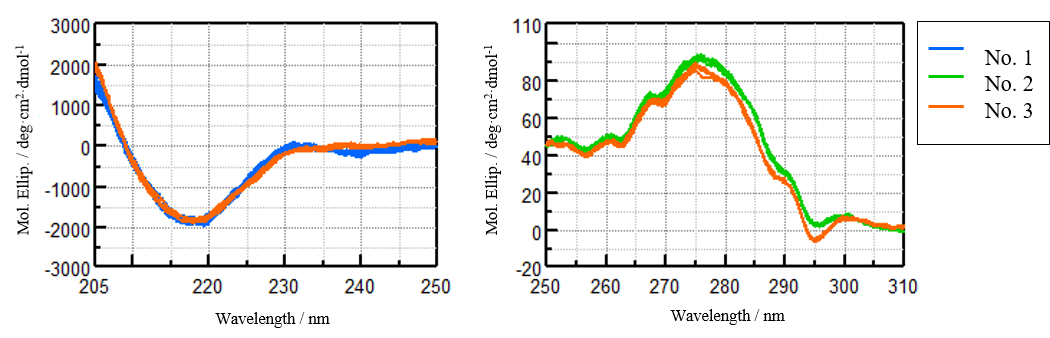
Figure 3. Far- UV (left) and near-UV (right) CD spectra of the VHH antibodies analyzed using the [Spectrum QC Compare Test] program.
The distance between the reference and sample CD spectra is shown in Figure 4 and the corresponding values are shown in Table 1. From Figure 4 it can be seen that weighting both the far-UV and the near-UV spectra increases the distance between reference and sample and the difference between them can be clearly confirmed.
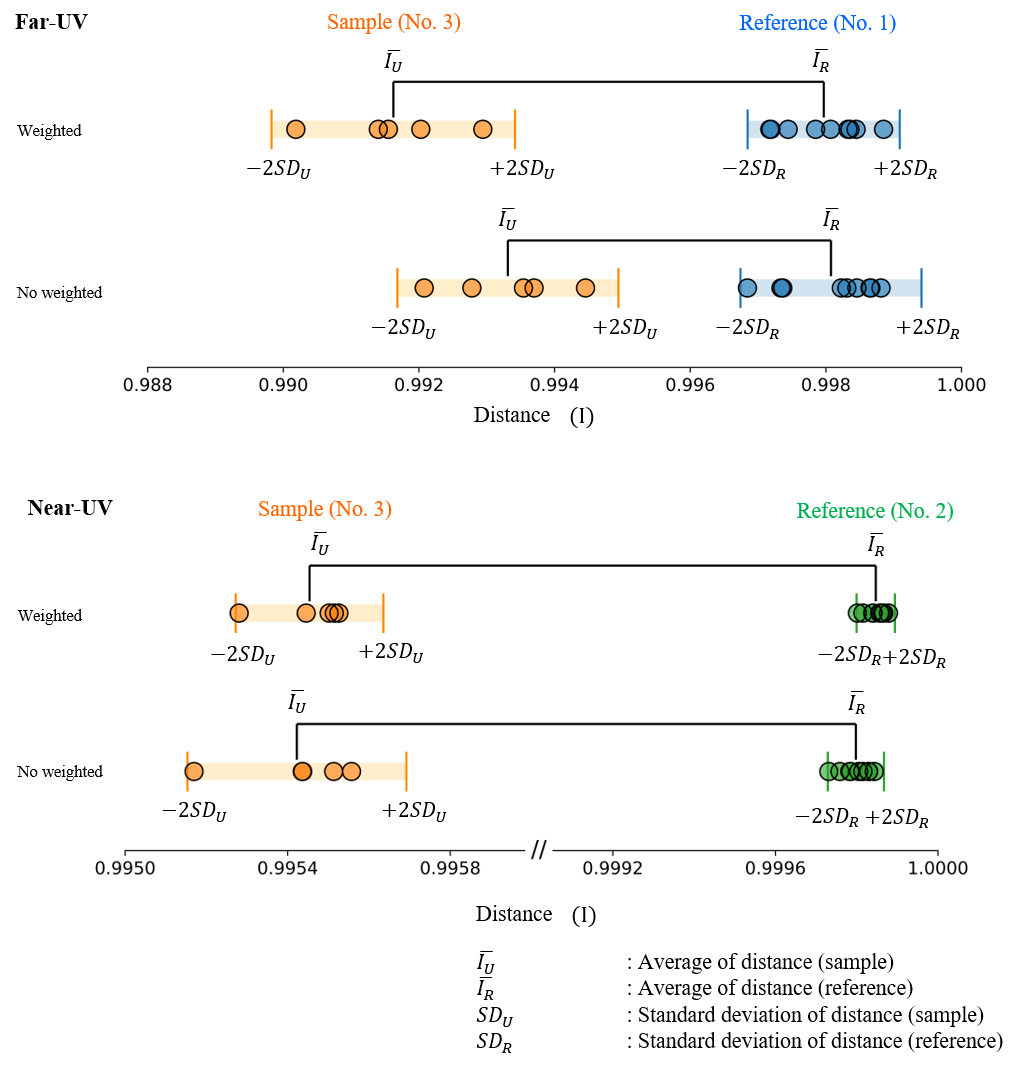
Figure 4. Calculated distances between the reference and sample CD spectra in the far-UV (top) and near-UV (bottom) using 2 standard deviations.
Table 1. Distances between the reference and sample far-UV (left) and near-UV (right) CD spectra.
Far-UV region
a) Reference (No. 1)
| Number of measurements | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 |
| Weighted | 0.9984 | 0.9983 | 0.9978 | 0.9972 | 0.9989 | 0.9984 | 0.9974 | 0.9972 | 0.9981 |
| No weighted | 0.9987 | 0.9985 | 0.9983 | 0.9968 | 0.9988 | 0.9987 | 0.9973 | 0.9974 | 0.9982 |
b)Sample (No. 3)
| Number of measurements | 1 | 2 | 3 | 4 | 5 |
| Weighted | 0.9916 | 0.9902 | 0.9929 | 0.9914 | 0.9920 |
| No weighted | 0.9935 | 0.9921 | 0.9945 | 0.9928 | 0.9937 |
Near-UV region
a) Reference (No. 2)
| Number of measurements | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 |
| Weighted | 0.99980 | 0.99988 | 0.99986 | 0.99986 | 0.99987 | 0.99984 | 0.99986 | 0.99984 | 0.99982 |
| No weighted | 0.99973 | 0.99984 | 0.99982 | 0.99983 | 0.99983 | 0.99978 | 0.99981 | 0.99979 | 0.99976 |
| Number of measurements | 1 | 2 | 3 | 4 | 5 |
| Weighted | 0.9955 | 0.9955 | 0.9955 | 0.9954 | 0.9953 |
| No weighted | 0.9956 | 0.9954 | 0.9954 | 0.9955 | 0.9952 |
In addition, the Z-scores are shown in Table 2. The Z-scores were all larger than 2 with and without weighting, indicating that there was a significant difference between the CD spectrum of the reference and sample both in the far- and near-UV.
Table 2. Calculated Z-scores from far-UV and near-UV CD spectra using [Spectrum QC Compare Test] program.
| Number of measurements | Far-UV | Near-UV | ||
| Weighted | No weighted | Weighted | No weighted | |
| 1 | 11.5 | 6.8 | 184 | 123 |
| 2 | 13.9 | 9.0 | 183 | 126 |
| 3 | 9.0 | 5.4 | 184 | 126 |
| 4 | 11.7 | 7.9 | 187 | 124 |
| 5 | 10.6 | 6.6 | 194 | 134 |
References
1) M. T. Gutierrez Lugo, U.S. Food and Drug Administration, Regulatory Consideration for the Characterization of HOS in Biotechnology Products, 5th International Symposium on Higher Order Structure of Protein Therapeutics 2016
2) W. F. W. 4th, J. P. Gabrielson, W. Al-Azzam, G. Chen, D. L. Davis, T. K. Das, D. B. Hayer, D. Houde, S. K, Technical Decision Making With Higher Order Structure Data: Perspectives on Higher Order Structure Characterization From the Biopharmaceutical Industry, J. Pharm. Sci., 105 (2016) 3465-3470
3) RePHAGEN Co., Ltd (https://rephagen.com/, May 20, 2020)
4) S. M. Kelly, N. C. Price, The Use of Circular Dichroism in the Investigation of Protein Structure and Function, Current Protein and Peptide Science, 1 (2000) 349-384.
Acknowledgements
Special thanks to RePHAGEN for providing VHH antibody samples
Many thanks to Dr. József Kardos at ELTE Eötvös Loránd University for scientific advice